You must have a 3 button mouse for this functionlity.
Basic Segmentation Test
- Read the Doxygen help page
- Load Analyze image NifTKData/Input/volunteers/16856/16856-002-1.img
- Make sure you are using the MIDAS "Drag and Drop Display" as your main editor window and not the MITK "Display".
- Drag and drop the image from the Datamanager to the Drag and Drop Display
- Check image does not appear obviously upside-down. If so, restart your application with environment variable NIFTK_DRC_ANALYZE=ON
- Open the MIDAS Morphological Editor. Window->Show View->MIDAS Morphological Editor
- Select the image in the Datamanager
- Click on "start/re-start segmentation" in the MIDAS Morphological Editor
- Set upper threshold to 139, lower threshold to 59 and axial cut-off to 96
- Click "accept"
- Set the number of erosions to 1
- Move to coronal slice 114, using the slice select slider at the top of the screen
- Click on the "Paintbrush tool", with a cursor size of 1.
- Use the connection breaker (middle mouse button) to remove (highlight in orange) the 3 pixels of the right optical chiasm (Figure 1.)
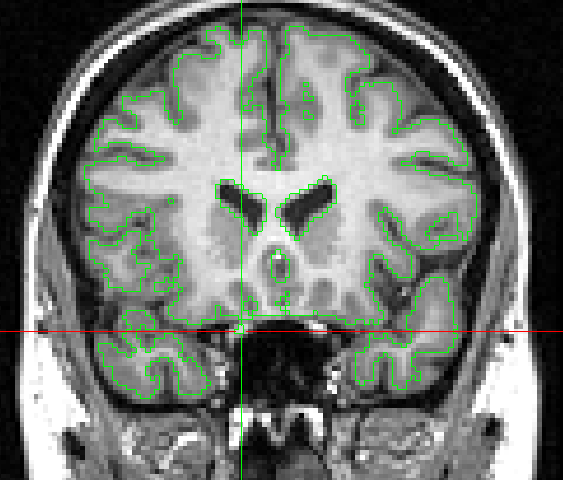
Figure 1. Slice 114, remove the right optical chiasm
- Ensure that the right optical chiasm has disappeared in slice 115 and 116
- Click "accept"
- Set the number of dilations to 2
- Move to coronal slice 119 using the mouse wheel to scroll
- Set the Paintbrush Tool to have a cursor size of 6.
- Use connection breaker to remove the right temporal lobe.
- Check the right temporal lobe has disappeared from slices 120+
- Click "reset"
- Check if you are at the Erosions tab and click "accept".
- Check the connection breaker is still present on slice 119
- Check if the number of dilations is still set to 2
- Check connection breaker has stopped the right temporal lobe expanding beyond slice 119
- Click "accept"
- Set the re-thresholding box size to 6
- Click "ok" to finish the segmentation
- Open the Image Statistics View. Window->Show View->Image Statistics
- In Datamanager, click on the image, and then the segmentation while holding the Ctrl (Window and Linux) or Cmd key (Mac). Make sure that you see "16856-002-1" in the "image:" line in the Image Statistics view and the chosen segmentation name after "mask:".
- In the Image Statistics View click "update"
- Check that the volume is 1178.33 (ml) +/- 0.25
- Check that the mean intensity value is 87.9 +/- 0.1
Additional Reliability Tests (Checking for not crashing)
- Restart the application.
- Load image 16856-002-1.img, drop the image into the Drag and Drop Display, close the Drag and Drop display.
- Right click on the image in the Datamanager, select Show In->Drag and Drop Display, check that the Drag and Drop Display is re-launched. At this stage the image will not be visible in the Drag and Drop Display. Drag and Drop the image from the Datamanager to the Drag and Drop Display again.
- Re-start the segmentation process above. After any stage select "cancel", which should abandon the segmentation.
- Re-start the segmentation process above. At any stage, try to exit the MIDAS Morphological Editor View. This should abandon the segmentation.
- Re-start the segmentation process above. At each tab, before clicking "accept", click "reset" which should return to the previous tab. You will then need to "accept" the previous tabs settings again to move forward. You should not be able to jump from one tab to the next and miss a step. If you can switch tabs, and move forward or backwards in the tab selector, the tab items should be disabled.
 1.8.8
1.8.8