Introduction
The MIDAS Morphological editor has been implemented within NiftyView using
- An MITK plugin:
NifTK/Code/Gui/MITK/Plugins/uk.ac.ucl.cmic/midasmorphologicalsegmentor
- Qt widgets, signals and slots.
- An abstract interface defining an ITK pipeline.
- ITK classes in:
NifTK/Code/Libs/ITK/Segmentation
- Unit tests for ITK classes in:
NifTK/Testing/Code/Libs/ITK/Segmentation
Class Overview
Figure 1 shows the class overview, and Figure 2 shows the initial view of the MIDAS Morphological Editor plugin.
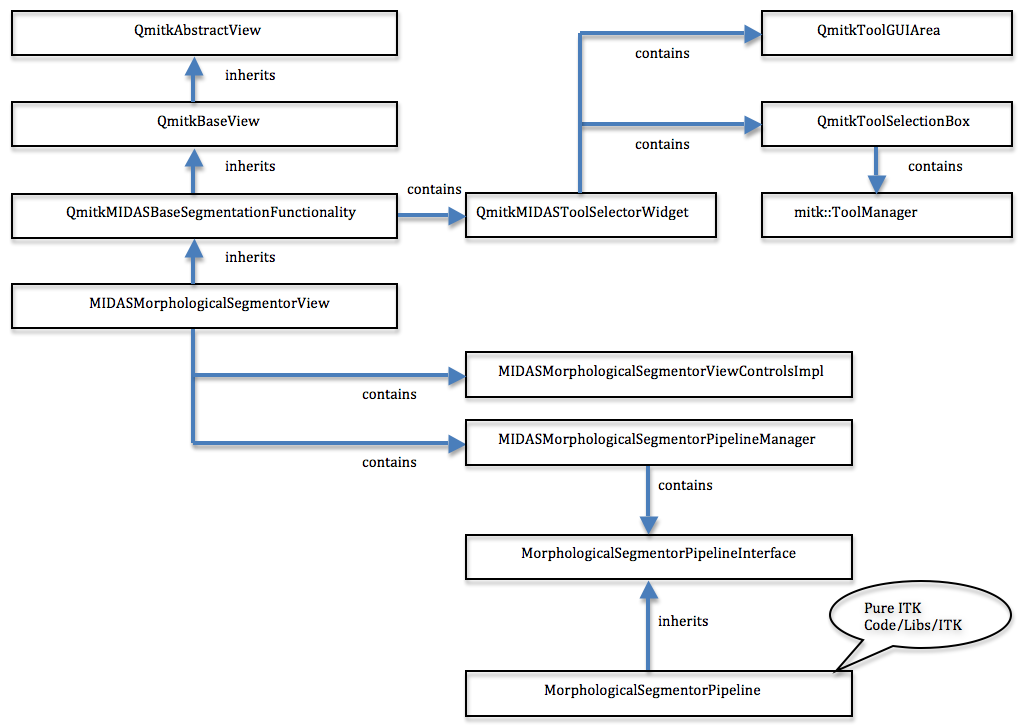
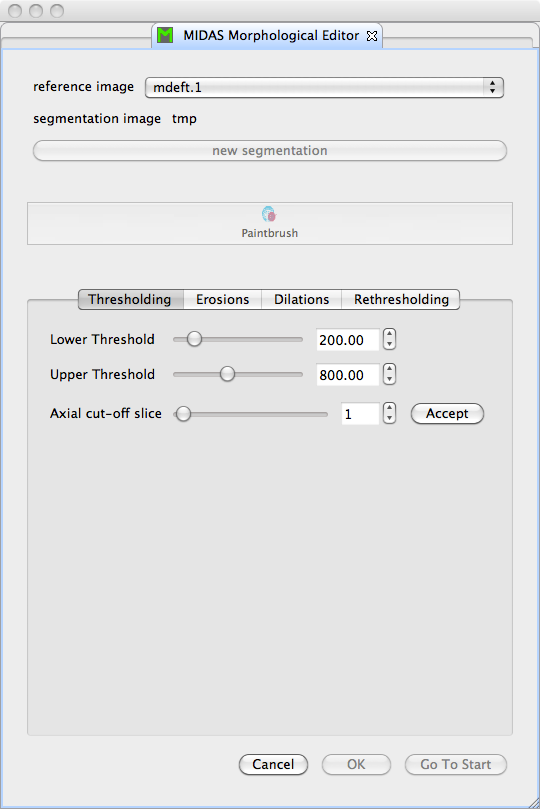
The MIDAS Morphological Editor is provided as an MITK plugin called MIDASMorphologicalSegmentorView derived from QAbstractView. The niftkBaseSegmentationView contains niftkMIDASImageAndSegmentationSelectorWidget providing the image selector and new segmentation button at the top of the plugin. niftkBaseSegmentationView also contains an MITK QmitkToolGUIArea and QmitkToolSelectionBox, and QmitkToolSelectionBox creates an mitk::ToolManager. These MITK classes are part of the MITK Segmentation Framework. To understand this plugin, you must read that paper.
The main GUI control is assembed in niftk::MorphologicalSegmentorView and comprises:
- a niftk::ImageAndSegmentationSelectorWidget contained in niftk::BaseSegmentatorView
- a niftk::ToolSelectorWidget contained in niftk::BaseSegmentatorView
- a niftk::MorphologicalSegmentorViewControlsImpl contained in niftk::MorphologicalSegmentorView
in that order. The GUI classes created in niftk::BaseSegmentatorView are also used in the niftk::GeneralSegmentorView, providing common functionality, look and feel for both plugins.
The GUI components use Qt signals and slots to call methods on niftk::MorphologicalSegmentorView. niftk::MorphologicalSegmentorView contains a niftk::MorphologicalSegmentorPipelineManager which itself contains a MorphologicalSegmentorPipelineInterface which is implemented by MorphologicalSegmentorPipeline using ITK. The whole plugin is essentially a Qt GUI, calling methods in niftk::MorphologicalSegmentorView which sets parameters on the ITK pipeline, which updates, and produces an output, which can be visualised using MITK and hence VTK as a binary image.
Pipeline Operation
The MIDAS Morphological Editor consists of 4 distinct phases: Thresholding, Erosions, Dilations and Rethresholding. This workflow can be seen in Figure 3, as 4 separate tabs that operate in sequence.
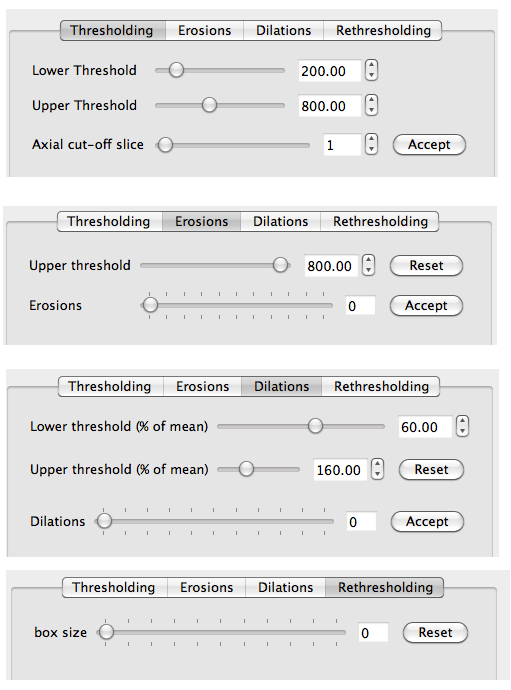
This workflow is directly mirrored by the ITK pipeline. Figure 4 shows the the ITK classes.
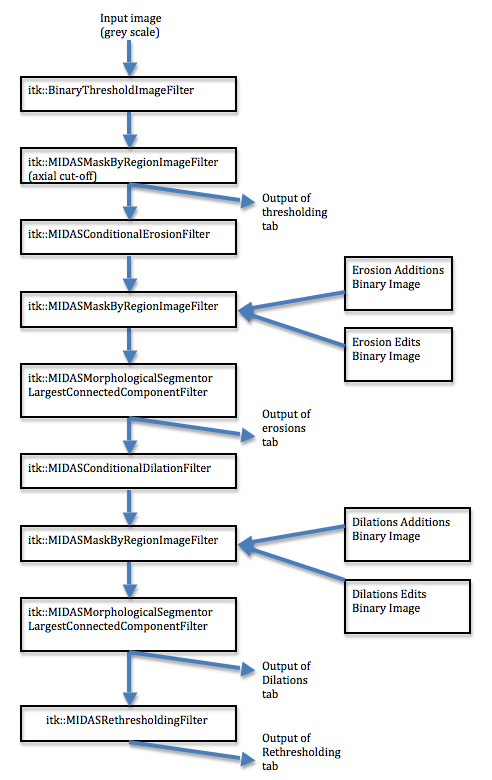
When the GUI is displaying the Thresholding tab, the pipeline is only updating itk::BinaryThresholdImageFilter and itk::MIDASMaskByRegionImageFilter. When the GUI is displaying the next 3 tabs, additional filters are added to the pipeline:
Pipeline Editing (Connection Breaker)
When the Erosions and Dilations tab are active, the user can edit the segmentation.
- Left mouse button = add to the segmentation - i.e. paint voxels in the Erosions Additions image or the Dilations Additions image which are added to the output in the corresponding itk::MIDASMaskByRegionImageFilter.
- Middle mouse button = break connections - this is highlighted in orange. This is achieved by painting voxels in the Erosions Edits or the Dilations Edits image, which are then used to subtract voxels out of the output of the corresponding itk::MIDASMaskByRegionImageFilter.
- Right mouse buttons = remove from the orange region used for connection breaking. This is achieved by painting voxels in the Erosions Edits or Dilations Edits image with a zero value, thereby erasing them. In effect this is an undo operation for the middle mouse button, leading to connections being un-broken.
The editing is implemented using the MITK Segmentation Framework. The mitk::ToolManager can manage a number of tools that can be used to edit segmentations. In the case of the MIDAS Morphological Editor, the mitk::ToolManager registers 4 working data images, these are the Additions image and the Edits image for both Erosions and Dilations. This is slightly different to how the framework normally operates. The framework normally operates with a single working data, and the output image that the user is interested in is that single working data. In our case, the working data are 4 intermediate image that are used to influence the output of the ITK pipeline.
Furthermore, the MITK undo/redo framework is used so that the niftk::PaintbrushTool can undo and redo the edits.
Conclusion
The MIDAS Morphological Editor is re-implemented as a Qt GUI calling an ITK pipeline, delivered within an MITK plugin. Future researchers can easily add filters to the pipeline, easily add unit tests for these filters, and easily provide additional editing tools to provide more complex editing functionality as desired. The plugin is not designed to be infinitely extensible as by design it is for a very specific task.
 1.8.8
1.8.8